Abstract
Background
The incidence and mortality of gastrointestinal (GI) tumors are high in China. Some studies suggest that the gut microbiota is related to the occurrence and development of tumors. At present, there are no prospective studies based on the correlation between gastrointestinal tumors and gut microbiota in the Chinese population. The objective of this report is to characterize the fecal microbiota in healthy control participants and patients with esophageal cancer, gastric cancer, and colorectal cancer.
Methods
Patients with locally advanced or metastatic esophageal, gastric, and colorectal cancer were enrolled, and healthy people were included as controls. 16S rRNA sequencing was used to analyze the characteristics of fecal microbiota. PICRUSt software was used for functional prediction.
Results
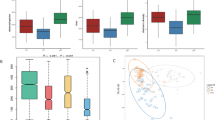
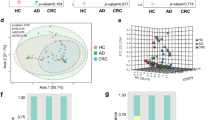
Significant differences in the composition and abundance of fecal microbiota were identified between gastrointestinal cancer patients (n = 130) and healthy controls (n = 147). The abundance of Faecalibacterium prausnitzii, Clostridium clostridioforme and Bifidobacterium adolescent in tumor groups were all significantly lower than in the control group (P < 0.05). The levels of Blautia producta and R. faecis in the gastric (n = 46) and colorectal cancer (n = 44) groups were significantly lower than those in the control group (P < 0.05). The level of Butyricicoccus pullicaecorum in the esophageal cancer (n = 40) and gastric cancer groups was significantly lower than that in the control group (P < 0.05). B. fragilis, Akkermansia muciniphila, Clostridium hathewayi and Alistipes finegoldii were overabundant in the different tumor groups compared with the control (P < 0.05). We observed significant differences in functional metabolism and cell biological function between the tumor and control groups (P < 0.05). Optimal microbial markers were identified on a random forest model and achieved an area under the curve of 85.59% between 130 GI cancer samples and 147 control samples. The respective AUC values were 86.89%, 97.11%, and 79.1% in detecting esophageal cancer, gastric cancer, and colorectal cancer.
Conclusions
Patients with esophageal or gastric cancers had similar features of fecal bacteria as those with colorectal cancer. The metabolic function of fecal bacteria in the gastrointestinal cancer patients and the healthy controls were different. The microbial signatures may potentially be applied to distinguish GI cancer patients from healthy people as a non-invasive diagnostic biomarker.






Similar content being viewed by others
Availability of data and materials
Please contact author for data requests.
Notes
Abbreviations
- GI:
-
Gastrointestinal
- EC:
-
Esophageal cancer
- GC:
-
Gastric cancer
- CRC:
-
Colorectal cancer
- rRNA:
-
Ribosomal RNA
- PD-L1:
-
Anti-programmed cell death ligand-1
- PD-1:
-
Anti-programmed cell death receptor-1
- PUMCH:
-
Peking Union Medical College Hospital
- ESR:
-
Erythrocyte sedimentation rate
- hsCRP:
-
Hypersensitive C-reactive protein
- ANA:
-
Antinuclear antibody
- OTU:
-
Operational taxonomic units
- PCoA:
-
Principal coordinate analysis
- ANOSIM:
-
Analysis of similarity
- LDA:
-
Linear discriminant analysis
- LEfSe:
-
Linear discriminant analysis effect size
- PICRUSt:
-
Phylogenetic Investigation of Communities by Reconstruction of Unobserved States
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- COG:
-
Clusters of Orthologous Groups of proteins
- ANOVA:
-
Analysis of variance
- FDR:
-
False discovery rate
- RF:
-
Random forest
- ROC:
-
Receiver operating characteristic
- NK:
-
Natural killer cell
- NF-kB:
-
Nuclear factor kappa beta
- STAT3:
-
Signal transducer and activator of transcription 3
- TNF-α:
-
Tumor necrosis factor-α
- COX-2:
-
Cyclooxygenase-2
- SCFA:
-
Short-chain fatty acids
- Treg:
-
Regulated T cells
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;67(1):7–30.
Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66(2):115–32.
Song M, Garrett WS, Chan AT. Nutrients, foods, and colorectal cancer prevention. Gastroenterology. 2015;148(6):1244-1260.e16.
Sender R, Fuchs S, Milo R. Revised estimates for the number of human and bacteria cells in the body. PLoS Biol. 2016;14(8):e1002533.
Jandhyala SM, Talukdar R, Subramanyam C, et al. Role of the normal gut microbiota. World J Gastroenterol. 2015;21(29):8787–803.
Garrett WS. Cancer and the microbiota. Science. 2015;348(6230):80–6.
Nougayrède JP, Homburg S, Taieb F, et al. Escherichia coli induces DNA double-strand breaks in eukaryotic cells. Science. 2006;313(5788):848–51.
Wang T, Cai G, Qiu Y, et al. Structural segregation of gut microbiota between colorectal cancer patients and healthy volunteers. ISME J. 2012;6(2):320–9.
Wu S, Rhee KJ, Albesiano E, et al. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat Med. 2009;15(9):1016–22.
Meng C, Bai C, Brown TD, et al. Human gut microbiota and gastrointestinal cancer. Genomics Proteom Bioinform. 2018;16(1):33–49.
Ren Z, Li A, Jiang J, et al. Gut microbiome analysis as a tool towards targeted non-invasive biomarkers for early hepatocellular carcinoma. Gut. 2019;68(6):1014–23.
Akshintala VS, Talukdar R, Singh VK, et al. The gut microbiome in pancreatic disease. Clin Gastroenterol Hepatol. 2019;17(2):290–5.
Geller LT, Barzily-Rokni M, Danino T, et al. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science. 2017;357(6356):1156–60.
Iida N, Dzutsev A, Stewart CA, et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science. 2013;342(6161):967–70.
Sivan A, Corrales L, Hubert N, et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science. 2015;350(6264):1084–9.
Gopalakrishnan V, Spencer CN, Nezi L, et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science. 2018;359(6371):97–103.
Routy B, Le Chatelier E, Derosa L, et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science. 2018;359(6371):91–7.
Kozich JJ, Westcott SL, Baxter NT, et al. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol. 2013;79(17):5112–20.
Magoč T, Salzberg SL. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics. 2011;27(21):2957–63.
Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 2010;26(19):2460–1.
Shannon CE. The mathematical theory of communication 1963. MD Comput. 1997;14(4):306–17.
Lozupone C, Lladser ME, Knights D, et al. UniFrac: an effective distance metric for microbial community comparison. ISME J. 2011;5(2):169–72.
Segata N, Izard J, Waldron L, et al. Metagenomic biomarker discovery and explanation. Genome Biol. 2011;12(6):R60.
Langille MG, Zaneveld J, Caporaso JG, et al. Predictive functional profifiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol. 2013;31(9):814–21.
Sommer F, Bäckhed F. The gut microbiota-masters of host development and physiology. Nat Rev Microbiol. 2013;11(4):227–38.
Wang Q, Garrity GM, Tiedje JM, et al. Naive Bayesian classifer for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol. 2007;73(16):5261–7.
Badal VD, Vaccariello ED, Murray ER, et al. The Gut Microbiome, Aging, and Longevity: A Systematic Review. Nutrients. 2020;12(12):3759.
Ahn J, Sinha R, Pei Z, et al. Human gut microbiome and risk for colorectal cancer. J Natl Cancer Inst. 2013;105(24):1907–11.
Lopez-Siles M, Martinez-Medina M, Surís-Valls R, et al. Changes in the abundance of Faecalibacterium prausnitzii Phylogroups I and II in the intestinal mucosa of inflammatory bowel disease and patients with colorectal cancer. Inflamm Bowel Dis. 2016;22(1):28–41.
Ohkusa T, Yoshida T, Sato N, et al. Commensal bacteria can enter colonic epithelial cells and induce proinflammatory cytokine secretion: a possible pathogenic mechanism of ulcerative colitis. J Med Microbiol. 2009;58(Pt 5):535–45.
Sinha R, Ahn J, Sampson JN, et al. Fecal microbiota, fecal metabolome, and colorectal cancer interrelations. PLoS ONE. 2016;11(3):e0152126.
Feng Q, Liang S, Jia H, et al. Gut microbiome development along the colorectal adenoma-carcinoma sequence. Nat Commun. 2015;6:6528.
Borges-Canha M, Portela-Cidade JP, Dinis-Ribeiro M, et al. Role of colonic microbiota in colorectal carcinogenesis: a systematic review. Rev Esp Enferm Dig. 2015;107(11):659–71.
Chen W, Liu F, Ling Z, et al. Human intestinal lumen and mucosa-associated microbiota in patients with colorectal cancer. PLoS ONE. 2012;7(6):e39743.
Chang S-C, Shen M-H, Liu C-Y, et al. A gut butyrate-producing bacterium Butyricicoccus pullicaecorumregulates short-chain fatty acid transporter and receptor to reduce the progression of 1,2-dimethylhydrazine-associated colorectal cancer. Oncol Lett. 2020;20(6):327.
Osaki T, Matsuki T, Asahara T, et al. Comparative analysis of gastric bacterial microbiota in Mongolian gerbils after long-term infection with Helicobacter pylori. Microb Pathog. 2012;53(1):12–8.
Wu N, Yang X, Zhang R, et al. Dysbiosis signature of fecal microbiota in colorectal cancer patients. Microb Ecol. 2013;66(2):462–70.
Wu S, Powell J, Mathioudakis N, et al. Bacteroides fragilis enterotoxin induces intestinal epithelial cell secretion of interleukin-8 through mitogen-activated protein kinases and a tyrosine kinase-regulated nuclear factor-kappaB pathway. Infect Immunity. 2004;72(10):5832–9.
Wu S, Morin PJ, Maouyo D, et al. Bacteroides fragilis enterotoxin induces c-Myc expression and cellular proliferation. Gastroenterology. 2003;124(2):392–400.
Weir TL, Manter DK, Sheflin AM, et al. Stool microbiome and metabolome differences between colorectal cancer patients and healthy adults. PLoS ONE. 2013;8(8):e70803.
Grivennikov SI, Wang K, Mucida D, et al. Adenoma-linked barrier defects and microbial products drive IL-23/IL-17-mediated tumour growth. Nature. 2012;491(7423):254–8.
Blackett KL, Siddhi SS, Cleary S, et al. Oesophageal bacterial biofilm changes in gastro-oesophageal reflux disease, Barrett’s and oesophageal carcinoma: association or causality? Aliment Pharmacol Ther. 2013;37(11):1084–92.
Aviles-Jimenez F, Vazquez-Jimenez F, Medrano-Guzman R, et al. Stomach microbiota composition varies between patients with non-atrophic gastritis and patients with intestinal type of gastric cancer. Sci Rep. 2014;4:4202.
Liang Q, Chiu J, Chen Y, et al. Fecal bacteria act as novel biomarkers for noninvasive diagnosis of colorectal cancer. Clin Cancer Res. 2017;23:2061–70.
Moschen AR, Gerner RR, Wang J, Reider SJ, et al. Lipocalin 2 protects from inflammation and tumorigenesis associated with gut microbiota alterations. Cell Host Microbe. 2016;19:455–69.
Agoff SN, Brentnall TA, Crispin DA, et al. The role of cyclooxygenase 2 in ulcerative colitis-associated neoplasia. Am J pathol. 2000;157(3):737–45.
IBD in EPIC Study Investigators, Tjonneland A, Overvad K, et al. Linoleic acid, a dietary n-6 polyunsaturated fatty acid, and the aetiology of ulcerative colitis: a nested case-control study within a European prospective cohort study. Gut. 2009;58(12):1606–11.
Kasala ER, Bodduluru LN, Barua CC, et al. Antioxidant and antitumor efficacy of Luteolin, a dietary flavone on benzo(a)pyrene-induced experimental lung carcinogenesis. Biomed Pharmacother. 2016;82:568–77.
Laviano A, Cascino A, Muscaritoli M, et al. Tumor-induced changes in host metabolism: a possible role for freetryptophan as a marker of neoplastic disease. Adv Exp Med Biol. 2003;527:363–6.
Zitvogel L, Ayyoub M, Routy B, et al. Microbiome and Anticancer Immunosurveillance. Cell. 2016;165(2):276–87.
Mima K, Sukawa Y, Nishihara R, et al. Fusobacterium nucleatum and T cells in colorectal carcinoma. JAMA Oncol. 2015;1(5):653–61.
Atarshi K, Tanoue T, Shima T, et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science. 2011;331(6015):337–41.
Hegazy AN, West NR, Stubbington MJT, et al. Circulating and tissue-resident CD4+ T cells with reactivity to intestinal microbiota are abundant in healthy individuals and function is altered during inflammation. Gastroenterology. 2017;153(5):1320–37.
Liang S, Xu L, Zhang D, et al. Effect of probiotics on small intestinal bacterial overgrowth in patients with gastric and colorectal cancer. Turk J Gastroenterol. 2016;27:227–32.
Funding
This work was supported by grants from the National Natural Science Foundation of China (No. 61435001), CAMS Innovation Fund for Medical Sciences (Nos. 2017-I2M-4-003, No.2016-I2M-1-001).
Author information
Authors and Affiliations
Contributions
NL designed the study, analyzed the data, and wrote the manuscript. CB, and LZ contributed to the data interpretation and the revision of the manuscript. NL, YG, and XL performed the sample collection. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This study was approved by the Ethics Committee of Peking Union Medical College Hospital.
Consent participate
All participants approved to participate, and written informed consent was obtained.
Consent for publication
All of the co-authors consented to publish this manuscript. A copy of the written consent is available for review.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, N., Bai, C., Zhao, L. et al. Characterization of the fecal microbiota in gastrointestinal cancer patients and healthy people. Clin Transl Oncol 24, 1134–1147 (2022). https://doi.org/10.1007/s12094-021-02754-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-021-02754-y