Abstract
Purpose
To identify a novel system for scoring intratumoral immune response that can improve prognosis and therapy decisions in early stage non-small cell lung cancer (NSCLC).
Methods/patients
Eighty-four completely resected stage I/II NSCLC without adjuvant therapy were classified by expression profiling using whole genome microarrays. An external cohort of 162 tumors was used to validate the results. Immune cells present in tumor microenvironment were evaluated semiquantitatively by CD20, CD79, CD3, CD8, CD4 and CD57 immunostaining. Univariate and multivariate analyses of variables associated with recurrence-free survival were performed.
Results
Initial molecular classification identified three clusters, one with significantly better RFS. A reduced two-subgroup classification and a 50-gene predictor were built and validated in an external dataset: high and low risk of recurrence patients (HR = 3.44; p = 0.001). Analysis of the predictor´s genes showed that the vast majority were related to a B/plasma cell immune response overexpressed in the low-risk subgroup. The predictor includes genes coding for unique B lineage-specific genes, functional elements or other genes that, although non-restricted to this lineage, have strong influence on B-cell homeostasis. Immunostains confirmed increased B-cells in the low-risk subgroup. Gene signature (p < 0.0001) and CD20 (p < 0.05) were predictors for RFS, while CD79 and K-RAS mutations showed a tendency.
Conclusions
Favorable prognosis in completely resected NSCLC is determined by a B-cell-mediated immune response. It can be differently scored by a 50-gene expression profile or by CD20 immunostaining. That prognosis information not reflected by traditional classifications may become a new tool for determining individualized adjuvant therapies.
Similar content being viewed by others
Introduction
Non-small cell lung cancer (NSCLC) represents 80 % of all lung cancers and constitutes a leading cause of death worldwide with 1.2 million new cases and over 1 million deaths per year [1]. Currently, TNM classification is the most important prognostic factor and guides treatment decisions [2]. Surgery with curative intent is the cornerstone for the management of early stages NSCLC, although a significant number of patients will relapse. The 5-year rates of local/regional recurrence reaches up to 16, 23, 37, 39 % for stage IA, IB, IIA or IIB patients [3]. Consequently, there is a pressing need for identifying patients at higher risk of relapse that could benefit from additional adjuvant therapy, but also from the potential identification of low-risk patients that could avoid the toxicity of this adjuvant treatment [4].
Gene expression microarrays have been extensively used to identify molecular signatures able to predict disease-free or overall survival. In NSCLC, prognostic signatures have been built in different tumor stages [5–17], by different analytical methods (supervised or unsupervised), for the most frequent histological subtypes: adenocarcinoma [5, 12, 13, 17] and Squamous Cell Carcinoma (SCC) [16].
Patients and methods
Patient selection
All patients diagnosed with NSCLC from 2001 to 2008 at Hospital Clinico San Carlos (HCSC), Madrid, Spain, with stages I/II (7th edition of the AJCC classification) were reviewed. Finally, 84 patients fulfilled the following criteria: (1) NSCLC tumors completely resected without close margins (R0) and with complete mediastinal lymph node dissection performed by the same surgical team, (2) no neoadjuvant/adjuvant treatment administered before/after surgery (at least until recurrence), (3) minimal follow-up if no recurrence was detected (24 months), (4) no evidence of other tumors at any time and (5) high-quality frozen tumor samples were available in the Biobank of our institution, with at least a 70 % tumor cell population and no extensive necrosis. Data regarding the clinicopathological variables were collected. No extracapsular extension was observed in the three stage IIB cases with positive nodes. The study protocol was approved by the Ethics Committee and the informed consent was obtained from each participant before enrollment in the study.
Tumor samples and RNA extraction
Total RNA was extracted using TRIZOL (Invitrogen, Carlsbad, CA) and a homogenizer (Ultraturrax T8-S8N-5G Rose Scientific Ltd., Canada) treated with DNAse using RNeasy Microkit (Qiagen GmbH, Germany). RNA quality was measured by Agilent Bioanalyzer 2100 (Agilent Technologies, Palo Alto CA, USA) and samples with a RIN (RNA Integrity Number) ≥7.5 were selected for the analysis.
Study of histopathological variables and mutational analysis of the tumors
Histopathological features such as tumor-infiltrating lymphocytes (TILs), acute inflammation, chronic inflammation or necrosis were assessed in tumor samples by two pathologists independently. TILs were considered positive when more than two lymphocytes per high power field (HPF) were identified within epithelial cell of the glandular tumoral component. Acute inflammation was considered positive when more than two polymorphonuclear cells (PMN) per HPF were identified. Chronic inflammation was classified as moderate or strong when lymphoplasmacytic cells were found in the tumoral stroma representing at least 10 or 20 %, respectively, of the total cell population; otherwise, it was classified as absent/weak. Necrosis was considered high when more than 5 % of the tumor showed necrotic areas.
Immunostains for immune cell markers (CD20, CD3, CD4, CD8, CD57, CD79) were performed on 5-μm thick sections obtained from paraffin-embedded tissue blocks, following our standard protocols, using DAKO ready-to-use antibodies, and DAKO Autostainer Plus System (Dako Cytomation, Carpinteria, CA, USA). Stained sections were reviewed by two pathologists semiquantitatively and the presence of positive cells throughout the whole section of the tumor was classified as weak or absent, moderate (10–30 cells/mm2) or strong (>30/mm2). Figure 4 shows representative cases.
K-RAS (codon 12 and 13), B-RAF (V600E) and EGFR (L858R in exon 21 and deletions in exon 19) were evaluated by PCR and capillary electrophoresis (see supplemental material)
Microarray profiling and data analysis
Gene expression profiles of 84 NSCLC were determined using two-color Agilent Microarrays (G4112F) as explained in Perez-Villamil et al. [18]. A pool of RNA obtained from 42 normal lung parenchyma samples was used as reference. As a quality control, SpikeIns were included. Procedures followed the recommendations of the manufacturer (manual G4140-90050.v5.7). Fluorescence was quantified and normalized (LOWESS) using Agilent scanner and Feature Extraction 10.7.3 software.
Data analysis was carried out using GeneSpring 11.5 (GS11.5) (Agilent) and BRB Array Tool 4.1 (BRB.4.1) developed by Dr. Richard Simon and BRB-ArrayTools Development Team. Flagged and absent spots were eliminated when present in more than 10 % of the samples. Spots representing the same gene (Entrez ID) were averaged. Resulting data set contained 84 tumor samples and 17,881 genes (g).
Tumor molecular classification
A list of 3,232 g was obtained after the exclusion of invariable genes and retaining only genes that had at least a threefold expression change from the median value in more than 10 % of samples. A second gene filtering excluded the genes differentially expressed between adenocarcinomas and SCCs (>1.5 fold-change and a p value ≤0.01 using an unpaired T Test with Benjamini and Hochberg correction (B & H). A gene list of 2,160 g was obtained. Tumors were classified following a two-step procedure: (a) Hierarchical clustering used to uncover the overall sample distribution; (b) K means applied to decrease intra-cluster heterogeneity and increase inter-cluster variability. Pearson centered correlation was applied to both techniques.
Development of the predictor and validation of results in an external cohort
To confirm the prognostic value of the molecular classification, a predictor was developed using the HCSC cohort (training) and applied to an external cohort (validation) that included 162 samples from the study by Roepman et al. [9]. Using Distance Weighted Discrimination-DWD-software [19], both data sets were combined in a single data set of 246 samples and 17,881 genes. Prediction analysis of microarrays (PAM) was used to generate the predictor [20].
Statistical analysis
We used SPSS 13.0 for Windows (SPSS Inc., Chicago, USA). Molecular subtype association with quantitative clinicopathological variables was analyzed by Kruskal–Wallis test. When necessary, post hoc tests were done using a Mann–Whitney U test. Chi Square/Fischer’s exact test assessed the association with qualitative parameters. All statistical analyses were two sided and considered significant when p value ≤0.05.
RFS was defined as the time from surgery to first confirmed relapse, otherwise the data were censored on the date of death or the last follow-up. Kaplan–Meier curves (K–M) and log-rank test assessed the probability of relapse-free survival for each molecular subtype and the other parameters evaluated. A Cox’s proportional hazards regression model was fitted to estimate survival distributions and hazard ratios. Finally, a multivariate Cox model was also built including variables with p < 0.1 from univariate analysis and clinically relevant prognostic variables.
Results
Histopathological variables description and prognostic evaluation
Table 1 describes the demographic features, histopathological variables and mutational analysis of our 84 NSCLC patients: 85.7 % males and 14.3 % females, mean age at diagnosis of 66.5 (±10.2) years old. Mean follow-up was 1,379 days with a 34.5 % recurrence rate. Seventy-one percent of the patients were stage I NSCLC and 28.6 % were stage II. Adenocarcinoma and SCC histopathological subtypes represented over 90 % of tumors. Presence of TILs and acute inflammation was observed in 63 and 43 % of the patients, respectively. Signs of chronic inflammation were present in almost all patients, but only 20 % of them were classified as moderate–strong inflammation. Mutational analysis revealed K-RAS alterations in 13 % of the patients, EGFR in 1.2 % and no B-RAF mutations were detected. No statistical association was found between RFS and histopathological subtype, tumor stage, chronic inflammation or any other clinicopathological parameter. Only K-RAS mutations showed a tendency towards a shorter RFS (p = 0.072) (Table 1).
Identification of three molecular subgroups and association with RFS: validation in the external cohort
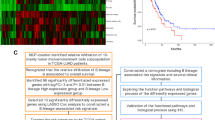
Microarray raw data are available in the ArrayExpress database (http://www.ebi.ac.uk/arrayexpress) under accession number E-MTAB-1790. Hierarchical clustering of the 84 samples using the 3,232 g distinguished two clusters (Supp. Fig. 2). Almost all SCC samples (97 %) were classified in one of the clusters (Cluster A) while 90 % adenocarcinoma cases were in the second cluster (Cluster B). No significant differences for RFS were found between Cluster A and B (p = 0.887). Since histological subtypes affected strongly tumor classification, we generated a new molecular classification after eliminating the genes differentially expressed between adenocarcinoma and SSC. The resulting list of 2,160 g was used to obtain a novel molecular classification of the 84 samples. Unsupervised analysis identified three clusters of approximately equal proportions (30, 25 and 29 samples respectively) (Supp. Fig. 3). Smoking status (p = 0.002), lymph node involvement (p = 0.041) and chronic inflammation (p = 0.002) showed a significant association with the molecular classification (Supp. Table 1). K–M curves revealed a strong association of the molecular classification with RFS (p = 0.004). Patients classified as Cluster-1 and -2 had shortened RFS than patients of Cluster-3 (Fig. 1). Cox regression model estimated a HR = 6.405 (95 % CI 1.8–22.3; p = 0.004) for Cluster-1 and a HR = 4.954 (95 % CI 1.4–17.8; p = 0.014) for Cluster-2 compared to Cluster-3. No significant differences were found between Cluster-1 and Cluster-2 (p = 0.526). Multivariate Cox model (Supp. Table 2) confirmed molecular classification as an independent prognostic factor after adjusting by tumor stage, CD20 and K-RAS status.
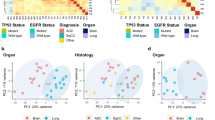
A 1,000 g predictor for the three molecular subgroups classification was developed from the list of 2,160 genes using the prediction analysis of microarrays technique (PAM) to reduce the number of genes and applied to the external cohort (n = 162) (Fig. 2). The cohort included 92 SCC, 58 AD, 10 LCC and 2 adenosquamous carcinomas. The rate of post-surgical recurrence was 34.5 % as a total of 56 patients developed tumor recurrence after surgery with curative intent.
Development of a low-risk predictor and validation in the external cohort
Since there were no differences in RFS between Clusters 1 and 2, we designed a simplified classification distinguishing only between high-risk (Cluster 1 + Cluster 2) and low-risk patients (Cluster 3). A 50 g predictor (Supp. Table 3) was generated and used to classify the validation dataset (n = 162) in the high- or the low-risk group. K–M curves showed a significant difference in RFS (p value = 0.001) (Fig. 3). Cox regression model confirmed the ability of our predictor to distinguish between high- and low-risk patients (HR = 3.44; 95 % CI 1.6–7.3; p = 0.001). Furthermore, our 50 g predictor was also able to recognize high- and low-risk patients within stages I and II, separately (Fig. 3a and b). HR of high-risk vs low-risk patients of 3.17 (95 % CI 1.2–8.3; p value = 0.018) for stage I and HR of 3.56 (95 % CI 1.1–12; p value = 0.041) for stage II.
Immunological nature of the 50-gene signature
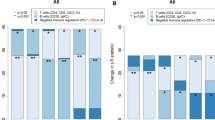
Analysis of the predictor’s 50 genes showed that the vast majority were related to a B/plasma cell immune response overexpressed in the low-risk subgroup. Gene Ontology (GO) term enrichment analysis revealed an over-representation of terms related to immune system process (GO:0002376; p = 3.47E−08), immune response (GO:0006955; p = 1.98E−08) and response to stimulus (GO:0050896; p = 0.023). Immunostaining for B-cell (CD20, CD79), T-cell (CD3, CD4, CD8) and NK-cell (CD57) markers confirmed an increased presence of B-cells in tumor microenvironment of the low-risk molecular subgroup while T and NK cells showed no significant differences among molecular subgroups (Table 1).
Immunostains also showed (Table 1; Fig. 4) that B-cell markers correlated with RFS: CD20 positivity (moderate or strong) (p = 0.05) and CD79 showed a tendency (p = 0.08). T-cell (CD3, CD4 and CD8) or NK-cell markers (CD57) showed no prognostic significance. Our 50-gene signature predicts RFS better than CD20 immunostaining and traditional TNM staging (Fig. 5).
Kaplan–Meier curves for RFS according to immunohistochemical markers in our series: a A strong presence of CD20 positive cells (×200). b Same tumor showing a moderate presence of CD3 positive cells (×200). c1 K–M curves for RFS for CD20 B-cell immunostaining: Strong (+++), moderate (++) or weak/absent (0/+), p value (Breslow) = 0.155. c2 K–M for CD20 moderate or strong (++/+++) vs absent/weak (0/+), p value = 0.05. d K–M for plasma cell marker CD79, p value = 0.309 (p = 0.08 when comparing strong +++ vs moderate/weak/absent, curves not shown). e K–M for CD3
Discussion
In this study, we describe the development of a 50-gene predictor that identifies a group of early stage completely resected NSCLC patients with a lower risk of recurrence. Molecular classification showing differences in tumor recurrence was obtained after eliminating the genes differentially expressed between adenocarcinoma and SCC. At early stages NSCLC, our classification was able to identify, independently of the TNM, one group of patients with lower likelihood of recurrence and two groups of patients with higher likelihood of recurrence. Since there were no differences in recurrence between the two clusters with higher risk, a simplified classification in two groups was established, one high and one low risk of recurrence groups of patients. A 50-gene low-risk predictor was built and validated in the external dataset. According to National Comprehensive Cancer Network (NCCN) guidelines, the TNM staging system and the extent of residual tumor after resection currently guides treatment decisions for stage I and II tumors. In the case of completely resected stage IA NSCLC, risk factors that signal the need for adjuvant chemotherapy include poor differentiation, vascular invasion, wedge resection, and minimal margins. However, the fact that disease relapse rates are as high as 30 %, even among stage IA patients [3]. Those conventional parameters for patient management, including tumor stage (IA, IB, IIA, IIB), and other clinicopathological variables showed a limited usefulness for predicting outcomes in our series of carefully selected R0 Stage I and II NSCLC, in agreement with other authors [4]. Only the presence of K-RAS mutations appears to be a poor prognostic indicator, in line with other authors [21]. Microarray study limitations due to a lack of appropriate experimental design have been recently referred by Subramanian et al. [22] Our study, to overcome these shortcomings [22, 23], carefully describes the patient inclusion criteria, patient characteristics, tissue handling and the statistical and analytical methods employed. Surgical procedures, inadequate mediastinal lymph node dissection, extracapsular spread, multiple positive hilar nodes, and close margins are variables that can influence recurrence beside TNM staging, not described in most previous expression profiling studies. Thus, our molecular classification includes samples belonging to different NSCLC subtypes making easier its application in other datasets and avoiding the shortcomings of predictors built with only one histopathological subtype pointed out by Houet et al. [13].
The identified 50 g predictor reflects by its composition the great importance of an immune response for the prognostic of lung cancer patients. The predictor´s 50 g were overexpressed in the low- compared to the high-risk group and the vast majority of them were, or might be, related to B/plasma cells. The predictor includes genes coding for unique B lineage-specific genes such as Ig molecules, B-cell receptor (CD79a) and B-cell lineage-specific marker (CD19) or for functional elements such as B-cell-specific transcriptional co-activator (POU2AF1) and marginal zone B1 cells (MZB1).Other genes, although non-restricted to this lineage, have strong influence on B-cell homeostasis such as TNFRSF17, SLAM7F, CXCL13 and IRF4. In contrast to the B/plasma cells, almost no lineage-specific genes for T-cells, NK cells or macrophages were differentially expressed between high- and low-risk patients. Although some of the predictor genes have traditionally been ascribed to T-cells, like Granzyme B, CD38, CD27 and another genes like Ig Fc receptors (FCRLA, FCRL5, FCRL2), they can also be expressed by B/plasma cells. For example, CD38 and CD27 are strongly upregulated in plasma cells/plasmablasts promoting their proliferation and survival [24], CD27 is also a marker of memory B-cells [25] and Pim-2 is an anti-apoptotic mediator, for the B-cell lineage as well [26, 27]. This is also the case for Granzyme B [28]. The finding that IGHA (alpha heavy chain) ranks in the first position of the 50 genes points out to infiltrating plasma cells migrated from blood, since most of circulating plasma cells are IgA + ve [24]. This may fit with the fact that EFNA4 has been shown to inhibit transmigration of B-cells through the vascular endothelium [28], since it is overexpressed in patients with worst prognosis (data not shown).
The only quite specific T-cell gene included in the 50 g predictor was LAX1. Its overexpression results in an inhibition of T-cell activation [29]. This fits with a recent study showing that “absence” of lung CD4 T-cells with an effector phenotype emerged as a predictor of favorable outcome in NSCLC [30]. Interestingly, Granzyme B, overexpressed in our cohort, can be secreted by B-cells differentiated into “cytotoxic” B-cells upon an incomplete T-cell help [31]. Previously, activated T-cells were linked to survival as detected by the gene expression of the quite T-cell-specific LCK [32], but this kinase can be expressed in many cancer cells [33] and even in B1 cells [34]. Immunostainings for different B, T or NK cells markers showed that T and NK cells can be detected in both high-risk and low-risk groups while B-cells in tumor microenvironment are more frequently found in the low-risk group (p < 0.05).Although the immune mechanisms with prognostic value have been related mainly to T-cells, the involvement of B/plasma cells has also been described in accordance with our results [35–37]. Ascierto et al. [35] described a 5-gene molecular signature of tumor-infiltrating immune B-cells that can predict a low risk of distant relapse in breast cancer patients regardless of ER, PR, or HER-2 status. The relationship of B/plasma cell infiltration and survival was also shown previously in medullary duct breast carcinomas [36] and related to an antigen-driven response directed to tumor-associated gangliosides [37].
Our gene signature can be used as an immunoscore for NSCLC. Gallon et al. [38] have recently postulated that the use of an immunoscoring, together with the TNM stage, is necessary to make prognostic more accurate in cancer. Different ways to quantify immune infiltration by light microscopy or immunohistochemistry have been used [39]. In our series, CD20 or CD79 immunostaining for B-cell or plasma cell, respectively, can be useful immunoscores, although the 50-gene signature is a more powerful tool to distinguish groups with different relapse-free survival probability (Fig. 4). Our predictor is able to identify patients with a worse prognosis that could benefit of additional treatment and patients with a good prognosis who may be candidates to avoid the toxic effects of adjuvant therapy. In conclusion, our results have the potential for improving early stage NSCLC patients management selecting those patients at lower risk of recurrence while opening new possibilities for immunotherapy and vaccine development.
References
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA Cancer J Clin. 2012;62(1):10–29.
Vallières E, Shepherd FA, Crowley J, Van Houtte P, Postmus PE, Carney D, et al. The IASLC Lung cancer staging project: proposals regarding the relevance of TNM in the pathologic staging of NSCLC in the forthcoming (seventh) edition of the TNM classification for lung cancer. J Thorac Oncol. 2009;4(9):1049–59.
Edge SB, Byrd DR, Compton CC, Fritz AG, Greene FL, Trotti A et al. eds.: AJCC cancer staging manual. 7th ed. New York: Springer; 2010. p. 253–704.
Agulló-Ortuño MT, López-Ríos F, Paz-Ares L. Lung cancer genomic signatures. J Thorac Oncol. 2010;5(10):1673–91.
Takeuchi T, Tomida S, Yatabe Y, Kosaka T, Osada H, Yanagisawa K, et al. Expression profile-defined classification of lung adenocarcinoma shows close relationship with underlying major genetic changes and clinicopathologic behaviors. J Clin Oncol. 2006;24:1679–88.
Sun Z, Wigle DA, Yang P. Non-overlapping and non-cell-type-specific gene expression signatures predict lung cancer survival. J Clin Oncol. 2008;26(6):877–83.
Zhu CQ, Ding K, Strumpf D, Weir BA, Meyerson M, Pennell N, et al. Prognostic and predictive gene signature for adjuvant chemotherapy in resected NSCLC. J Clin Oncol. 2010;28(29):4417–24.
Lu Y, Lemon W, Liu P-Y, Yi Y, Morrison C, Yang P, et al. A gene expression signature predicts survival of patients with stage I NSCLC. PLoS Med. 2006;3(12):e467.
Roepman P, Jassem J, Smit EF, Muley T, Niklinski J, van de Velde T, et al. An immune response enriched 72-gene prognostic profile for early-stage NSCLC. Clin Cancer Res. 2009;15(1):284–90.
Zhu CQ, Pintilie M, John T, Strumpf D, Shepherd FA, Der SD, et al. Understanding prognostic gene expression signatures in lung cancer. Clin Lung Cancer. 2009;10(5):331–40.
Lau SK, Boutros PC, Pintilie M, Blackhall FH, Zhu CQ, Strumpf D, et al. Three-gene prognostic classifier for early-stage NSCLC. J Clin Oncol. 2007;25:5562–9.
Neumann J, Feuerhake F, Kayser G, Wiech T, Aumann K, Passlick B, et al. Gene expression profiles of lung adenocarcinoma linked to histopathological grading and survival but not to EGF-R status: a microarray study. BMC Cancer. 2010;10:77.
Hou J, Aerts J, den Hamer B, van Ijcken W, den Bakker M, Riegmanet P et al. Gene expression-based classification of NSCLC and survival prediction. PloS One 2010; 5(4): e10312.
Bryant CM, Albertus DL, Kim S, Chen G, Brambilla C, Guedj M et al. Clinically relevant characterization of lung adenocarcinoma subtypes based on cellular pathways: an international validation study. PLoS One 2010;22; 5(7):e11712.
Wan YW, Sabbagh E, Raese R, Qian Y, Luo D, Denvir J et al. Hybrid models identified a 12-gene signature for lung cancer prognosis and chemoresponse prediction. PLoS One 2010;17;5(8):e12222.
Zhu CQ, Strumpf D, Li CY, Li Q, Liu N, Der S, et al. Prognostic gene expression signature for SCC of lung. Clin Cancer Res. 2010;16:5038.
Kratz JR, He J, Van Den Eeden SK, Zhu ZH, Gao W, Pham PT, et al. A practical molecular assay to predict survival in resected non-squamous, NSCLC: development and international validation studies. Lancet. 2012;379:823–32.
Perez-Villamil B, Romera-Lopez A, Hernandez-Prieto S, Lopez Campos G, Calles A, Sanz-Ortega J, et al. Colon cancer molecular subtypes identified by expression profiling and associated to stroma, mucinous type and different clinical behavior. BMC Cancer. 2012;12:260.
Benito M, Parker J, Du Q, Wu J, Xiang D, Perou CM, et al. Adjustment of systematic microarray data bases. Bioinformatics. 2003;20(1):105–14.
Tibshirani R, Hastie T, Narasimhan B, Chu G. Diagnosis of multiple cancer types by shrunken centroids of gene expression. Proc Natl Acad Sci USA. 2002;99(10):6567–72.
Mascaux C, Iannino N, Martin B, Paesmans M, Berghmans T, Dusart M et al. The role of RAS oncogene in survival of patients with lung cancer: a systematic review of the literature with meta-analysis. Br J Cancer 2005;92:131–135.
Subramanian J, Simon R. Gene expression-based prognostic signatures in lung cancer: ready for clinical use? J Natl Cancer Inst. 2010;102:464–74.
Dupuy A, Simon RM. Critical review of published microarray studies for cancer outcome and guidelines on statistical analysis and reporting. J Natl Cancer Inst. 2007;99(2):147–57.
Perez-Andres M, Paiva B, Nieto WG, Caraux J, Schmitz CA, Almeida J, et al. Human peripheral blood B-cell compartments: a crossroad in B-cell traffic. Cytometry B Clin Cytom. 2010;78B(Suppl. 1):S47–60.
Tangye SG, Good KL. Human IgM+ CD27+ B cells: memory B cells or “memory” B cells? J Immunol. 2007;179(1):13–9.
White E. The pims and outs of survival signaling: role for the Pim-2 protein kinase in the suppression of apoptosis by cytokines. Genes Dev. 2003;17(15):1813–6.
Asano J, Nakano A, Oda A, Amou H, Hiasa M, Takeuchi K et al. The serine/threonine kinase Pim-2 is a novel anti-apoptotic mediator in myeloma cells. Leukemia 2011;25(7):1182–8.
Trinidad EM, Ballesteros M, Zuloaga J, Zapata A, Alonso-Colmenar LM. An impaired transendothelial migration potential of chronic lymphocytic leukemia (CLL) cells can be linked to ephrin-A4 expression. Blood. 2009;114(24):5081–90.
Shapiro MJ, Nguyen CT, Aghajanian H, Zhang W, Shapiro VS. Negative regulation of TCR signaling by linker for activation of X cells via phosphotyrosine-dependent and -independent mechanisms. J Immunology. 2008;181:7055–61.
Zikos AD, Donnenberg RJ, Landreneau JD, Luketich JD, Donnenberg VS. Lung T-cell subset composition at the time of surgical resection is a prognostic indicator in NSCLC. Cancer Immunol Immunother. 2011;60:819–27.
Hagn M, Sontheimer K, Dahlke K, Brueggemann S, Kaltenmeier C, Beyer T, et al. Human B cells differentiate into granzyme B-secreting cytotoxic B lymphocytes upon incomplete T-cell help. Immunol Cell Biol. 2012;90(4):457–67.
Chen H-Y, Yu S-L, Chen C-H, Chang GC, Chen CY, Yuan A, et al. A five gene signature and clinical outcome in NSCLC. N Engl J Med. 2007;356:11–20.
Mahabeleshwar GH, Das R, Kundu GC. Tyrosine kinase, p56lck-induced cell motility, and urokinase-type plasminogen activator secretion involve activation of EGFR/extracellular signal regulated kinase pathways. J BiolChem. 2004;279:9733–42.
Majolini MB, D’Elios MM, Galieni P, Boncristiano F, Lauria G, Del Prete JL, et al. Expression of the T-cell-specific tyrosine kinase lck in normal B-1 cells and in chronic lymphocytic leukemia B cells. Blood. 1998;91:3390–6.
Ascierto ML, Kmieciak M, Idowu MO, Manjili R, Zhao Y, Grimes M, et al. A signature of immune function genes associated with recurrence-free survival in breast cancer patients. Breast Cancer Res Treat. 2012;131(3):871–80.
Coronella JA, Telleman P, Kingsbury GA, Truong TD, Hays S, Junghans RP, et al. Evidence for an antigen-driven humoral immune response in medullary ductal breast cancer. Cancer Res. 2001;61(21):7889–99.
Kotlan B, Simsa P, Teillaud JL, Fridman WH, Toth J, McKnight M, et al. Novel ganglioside antigen identified by B cells in human medullary breast carcinomas: the proof of principle concerning the tumor-infiltrating B lymphocytes. J Immunol. 2005;175(4):2278–85.
Galon J, Pagès F, Marincola FM, Thurin M, Trinchieri G, Fox BA, et al. The immune score as a new possible approach for the classification of cancer. J Transl Med. 2012;10:1–4.
Suzuki K, Kachala SS, Kadota K, Shen R, Mo Q, Beer DG, et al. Prognostic immune markers in NSCLC. Clin Cancer Res. 2011;17(16):5247–56.
Hernando F, Sanz J, Jarabo JR, et al. A new genetic signature defining two prognostic groups among patients with completely resected early non-small cell lung cancer. J Thor Oncol. 2012;7(6):S59.
Sanz J, Hernandez S, Hernando F, Larriba JLG, Puente J, Ferrer M, et al. An intratumoral B-cell immune response determines favorable prognosis for early stage lung cancer and can be assessed by different immunoscores. Modern Pathol. 2013;26:466A.
Acknowledgments
We thank Dr. Richard Simon for his kind suggestions. We thank Dr. Paul Roepman and Nico van Zandwijk for gently providing us with their microarray and patient’s data. We thank Biobank of IDISSC as a sample bio-repository. This study has been funded by FIS, INMUNOTHERCAN-CM, S2010/BMD-2326 and RTICC-Instituto Salud Carlos III ref 06/0020/0021.
Conflict of interest
All authors state that there is no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hernández-Prieto, S., Romera, A., Ferrer, M. et al. A 50-gene signature is a novel scoring system for tumor-infiltrating immune cells with strong correlation with clinical outcome of stage I/II non-small cell lung cancer. Clin Transl Oncol 17, 330–338 (2015). https://doi.org/10.1007/s12094-014-1235-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-014-1235-1