Abstract
Background
Colorectal cancer is one of the most common malignancies. With continuous exploration of the interaction between tumor cells and the immune system, tumor immunotherapy has become a revolution. However, CRC remains one of the less effective tumors for immunotherapy. The tumor microenvironment plays an important role in tumorigenesis and progression. The aim of this study is to explore tumor microenvironment-related genes that can predict the prognosis of colorectal adenocarcinoma, and also to provide new ideas for the mechanism of tumor development as well as immunotherapy.
Methods
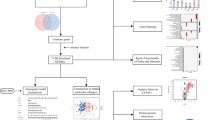
After estimating Stromalscore and Immunescore of colorectal adenocarcinoma tumor samples according to RNA-Seq expression data downloaded from TCGA, we screened for TME-related differential genes. We filtered prognosis-related core genes by constructing protein–protein interaction networks and making one-factor cox analysis for prognosis. Finally, the relative content of 22 immune cells in tumor tissues was evaluated, and then immune cells associated with core genes were identified.
Results
We screened 773 differential genes related to the TME. Then we identified C3 as a core gene associated with prognosis. Single gene analysis showed that C3 expression was significantly higher in tumor tissues than in normal tissues (p < 0.001). High C3 expression was associated with lower overall survival (p = 0.046). Tumor immune cell analysis showed that mast cells resting, mast cells activated, T cells CD4 memory activated, eosinophils, and macrophages M0 were C3-associated immune cells.
Conclusions
C3 has potential as a biomarker for colorectal adenocarcinoma and could provide new research ideas for the diagnosis and treatment of colorectal adenocarcinoma, especially for immunotherapy.






Similar content being viewed by others
Data availability
All relevant data are within the paper.
Code availability
All relevant codes are within the paper.
References
Bray F, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424.
Kasi PM, et al. Rising proportion of young individuals with rectal and colon cancer. Clin Colorectal Cancer. 2019;18(1):e87–95.
Wolf AMD, et al. Colorectal cancer screening for average-risk adults: 2018 guideline update from the American Cancer Society. CA Cancer J Clin. 2018;68(4):250–81.
Ait Ouakrim D, et al. Trends in colorectal cancer mortality in Europe: retrospective analysis of the WHO mortality database. BMJ. 2015;351:h4970.
Kirkegaard H, et al. Association of adherence to lifestyle recommendations and risk of colorectal cancer: a prospective Danish cohort study. BMJ. 2010;341:c5504.
Henrikson NB, et al. Family history and the natural history of colorectal cancer: systematic review. Genet Med. 2015;17(9):702–12.
Nguyen LH, Goel A, Chung DC. Pathways of colorectal carcinogenesis. Gastroenterology. 2020;158(2):291–302.
Harada S, Morlote D. Molecular pathology of colorectal cancer. Adv Anat Pathol. 2020;27(1):20–26.
Pino MS, Chung DC. Microsatellite instability in the management of colorectal cancer. Expert Rev Gastroenterol Hepatol. 2011;5(3):385–99.
Bae JM, Kim JH, Kang GH. Epigenetic alterations in colorectal cancer: the CpG island methylator phenotype. Histol Histopathol. 2013;28(5):585–95.
Guinney J, et al. The consensus molecular subtypes of colorectal cancer. Nat Med. 2015;21(11):1350–6.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–74.
Kalluri R, Zeisberg M. Fibroblasts in cancer. Nat Rev Cancer. 2006;6(5):392–401.
Straussman R, et al. Tumour micro-environment elicits innate resistance to RAF inhibitors through HGF secretion. Nature. 2012;487(7408):500-U118.
Sato E, et al. Intraepithelial CD8(+) tumor-infiltrating lymphocytes and a high CD8(+)/regulatory T cell ratio are associated with favorable prognosis in ovarian cancer. Proc Natl Acad Sci USA. 2005;102(51):18538–43.
Mlecnik B, et al. Histopathologic-based prognostic factors of colorectal cancers are associated with the state of the local immune reaction. J Clin Oncol. 2011;29(6):610–8.
Dekker E, et al. Colorectal cancer. Lancet. 2019;394(10207):1467–80.
Galon J, et al. Cancer classification using the Immunoscore: a worldwide task force. J Transl Med. 2012;10:205.
Fukunaga A, et al. CD8(+) tumor-infiltrating lymphocytes together with CD4(+) tumor-infiltrating lymphocytes and dendritic cells improve the prognosis of patients with pancreatic adenocarcinoma. Pancreas. 2004;28(1):E26–31.
Tomsova M, et al. Prognostic significance of CD3+ tumor-infiltrating lymphocytes in ovarian carcinoma. Gynecol Oncol. 2008;108(2):415–20.
Kryczek I, et al. Phenotype, distribution, generation, and functional and clinical relevance of Th17 cells in the human tumor environments. Blood. 2009;114(6):1141–9.
Sharma P, et al. CD8 tumor-infiltrating lymphocytes are predictive of survival in muscle-invasive urothelial carcinoma. Proc Natl Acad Sci USA. 2007;104(10):3967–72.
Pages F, et al. In situ cytotoxic and memory T cells predict outcome in patients with early-stage colorectal cancer. J Clin Oncol. 2009;27(35):5944–51.
Ogino S, et al. Lymphocytic reaction to colorectal cancer is associated with longer survival, independent of lymph node count, microsatellite instability, and CpG island methylator phenotype. Clin Cancer Res. 2009;15(20):6412–20.
Mlecnik B, et al. Integrative analyses of colorectal cancer show immunoscore is a stronger predictor of patient survival than microsatellite instability. Immunity. 2016;44(3):698–711.
Le YY, et al. Chemokines and chemokine receptors: their manifold roles in homeostasis and disease. Cell Mol Immunol. 2004;1(2):95–104.
Tanaka A, Sakaguchi S. Regulatory T cells in cancer immunotherapy. Cell Res. 2017;27(1):109–18.
Plitas G, et al. Regulatory T cells exhibit distinct features in human breast cancer. Immunity. 2016;45(5):1122–34.
Peng D, et al. Epigenetic silencing of TH1-type chemokines shapes tumour immunity and immunotherapy. Nature 2015;527(7577):249–53.
Karin N, Wildbaum G. The role of chemokines in shaping the balance between CD4(+) T cell subsets and its therapeutic implications in autoimmune and cancer diseases. Front Immunol. 2015;6:609.
Wanninger J, et al. Adiponectin-stimulated CXCL8 release in primary human hepatocytes is regulated by ERK1/ERK2, p38 MAPK, NF-kappa B, and STAT3 signaling pathways. Am J Physiol Gastrointest Liver Physiol. 2009;297(3):611–8.
Nastase A, et al. Expression of interleukine-8 as an independent prognostic factor for sporadic colon cancer dissemination. J Med Life. 2014;7(2):215–9.
Zumwalt TJ, et al. Active secretion of CXCL10 and CCL5 from colorectal cancer microenvironments associates with GranzymeB(+) CD8(+) T-cell infiltration. Oncotarget 2015;6(5):2981–91.
Colvin RA, et al. Intracellular domains of CXCR3 that mediate CXCL9, CXCL10, and CXCL11 function. J Biol Chem. 2004;279(29):30219–27.
Arenberg DA, et al. Improved survival in tumor-bearing SCID mice treated with interferon-gamma-inducible protein 10 (IP-10/CXCL10). Cancer Immunol Immunother. 2001;50(10):533–8.
Toiyama Y, et al. Evaluation of CXCL10 as a novel serum marker for predicting liver metastasis and prognosis in colorectal cancer. Int J Oncol. 2012;40(2):560–6.
Boguslawska J, et al. Expression of genes involved in cellular adhesion and extracellular matrix remodeling correlates with poor survival of patients with renal cancer. J Urol. 2016;195(6):1892–902.
Lyu T, et al. SMYD3 promotes implant metastasis of ovarian cancer via H3K4 trimethylation of integrin promoters. Int J Cancer. 2020;146(6):1553–67.
Sato T, et al. Interleukin 10 production by human melanoma. Clin Cancer Res. 1996;2(8):1383–90.
Li C, et al. TLR4 signaling pathway in mouse Lewis lung cancer cells promotes the expression of TGF-beta1 and IL-10 and tumor cells migration. Biomed Mater Eng. 2014;24(1):869–75.
Boulland ML, et al. Human interleukin-10 expression in T/natural killer-cell lymphomas: association with anaplastic large cell lymphomas and nasal natural killer-cell lymphomas. Am J Pathol. 1998;153(4):1229–37.
Kundu N, et al. Antimetastatic and antitumor activities of interleukin 10 in a murine model of breast cancer. J Natl Cancer Inst. 1996;88(8):536–41.
Zheng LM, et al. Interleukin-10 inhibits tumor metastasis through an NK cell-dependent mechanism. J Exp Med. 1996;184(2):579–84.
Mannino MH, et al. The paradoxical role of IL-10 in immunity and cancer. Cancer Lett. 2015;367(2):103–7.
Li BS, et al. Predictive value of IL-18 and IL-10 in the prognosis of patients with colorectal cancer. Oncol Lett. 2019;18(1):713–9.
Ricklin D, et al. Complement: a key system for immune surveillance and homeostasis. Nat Immunol. 2010;11(9):785–97.
Ricklin D, Reis ES, Lambris JD. Complement in disease: a defence system turning offensive. Nat Rev Nephrol. 2016;12(7):383–401.
Kwak JW, et al. Complement activation via a C3a receptor pathway alters CD4(+) T lymphocytes and mediates lung cancer progression. Can Res. 2018;78(1):143–56.
Janelle V, et al. Transient complement inhibition promotes a tumor-specific immune response through the implication of natural killer cells. Cancer Immunol Res. 2014;2(3):200–6.
Cho MS, et al. Complement component 3 is regulated by TWIST1 and mediates epithelial-mesenchymal transition. J Immunol. 2016;196(3):1412–8.
Downs-Canner S, et al. Complement inhibition: a novel form of immunotherapy for colon cancer. Ann Surg Oncol. 2016;23(2):655–62.
Roumenina LT, et al. Context-dependent roles of complement in cancer. Nat Rev Cancer. 2019;19(12):698–715.
Habermann JK, et al. Increased serum levels of complement C3a anaphylatoxin indicate the presence of colorectal tumors. Gastroenterology. 2006;131(4):1020–9.
Mehrabani D, et al. Clinical significance of serum vascular endothelial growth factor and complement 3a levels in patients with colorectal cancer in southern Iran. Asian Pac J Cancer Prev. 2014;15(22):9713–7.
Fridman WH, et al. The immune contexture in cancer prognosis and treatment. Nat Rev Clin Oncol. 2017;14(12):717–34.
Bonavita E, et al. PTX3 is an extrinsic oncosuppressor regulating complement-dependent inflammation in cancer. Cell. 2015;160(4):700–14.
Crivellato E, Ribatti D. The mast cell: an evolutionary perspective. Biol Rev. 2010;85(2):347–60.
Reichman H, et al. Activated eosinophils exert antitumorigenic activities in colorectal cancer. Cancer Immunol Res. 2019;7(3):388–400.
Gangwar RS, et al. Mast cell and eosinophil surface receptors as targets for anti-allergic therapy. Pharmacol Ther. 2017;170:37–63.
Sallusto F, Geginat J, Lanzavecchia A. Central memory and effector memory T cell subsets: function, generation, and maintenance. Annu Rev Immunol. 2004;22:745–63.
Fridman WH, et al. The immune contexture in human tumours: impact on clinical outcome. Nat Rev Cancer. 2012;12(4):298–306.
Funding
Not applicable.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All the authors declared that they have no conflict of interest.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, Y., Wang, X. Tumor microenvironment-associated gene C3 can predict the prognosis of colorectal adenocarcinoma: a study based on TCGA. Clin Transl Oncol 23, 1923–1933 (2021). https://doi.org/10.1007/s12094-021-02602-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-021-02602-z